fig4
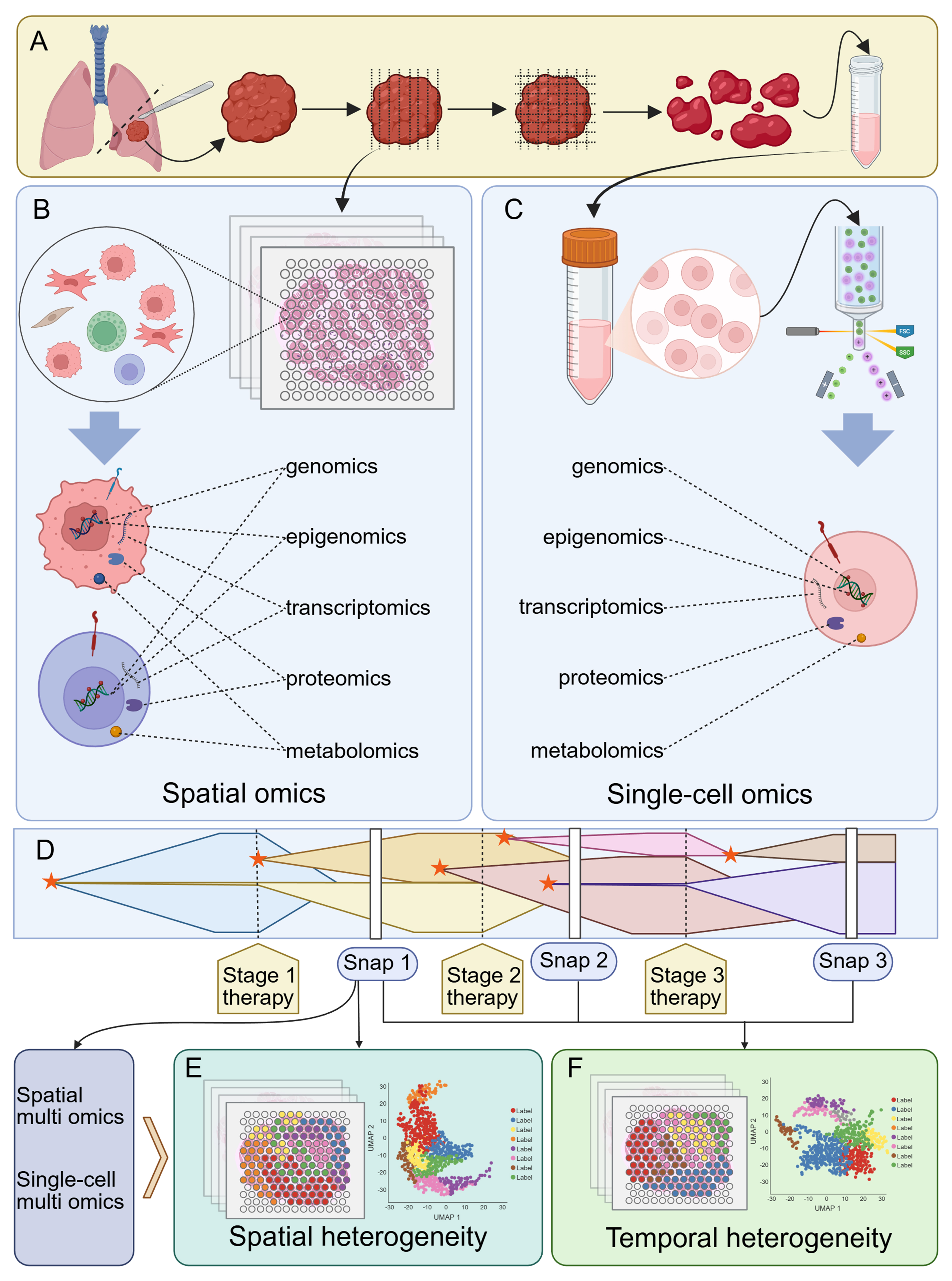
Figure 4. Investigating tumor resistance mechanisms associated with heterogeneity using single-cell and spatial multi-omics technologies. (A) Tumor tissue processing workflow. Tumor tissue obtained via surgical resection or other methods can be processed into slices for spatial multi-omics analysis, whereas tissue lysates yielding single-cell suspensions can be used for subsequent single-cell multi-omics analysis; (B) Spatial multi-omics analysis. Tissue slices are segmented into multiple spatial spots, with all cells within each spot undergoing comprehensive multi-omics profiling; (C) Single-cell multi-omics analysis. Single-cell suspensions undergo multi-step sorting and processing, ultimately enabling cell-level multi-omics profiling; (D) Diagram illustrating tumor heterogeneity during development. Tumor evolution involves multiple subclonal formation processes, with different subclones coexisting and undergoing selection and evolution under drug pressure or other survival stresses. Throughout this process, tumor tissue can be sampled at multiple time points (snaps) and analyzed using spatial or single-cell multi-omics approaches. A single snap data point addresses spatial tumor heterogeneity (E), whereas combined analysis of multiple data points addresses temporal tumor heterogeneity (F). The asterisks indicate subclonal formation. Created in BioRender. Da, E. (2026) https://BioRender.com/5zem2e5.